Paul Thompson
West Group
EMoE: Eigenbasis-Guided Routing for Mixture-of-Experts
Jan 17, 2026Abstract:The relentless scaling of deep learning models has led to unsustainable computational demands, positioning Mixture-of-Experts (MoE) architectures as a promising path towards greater efficiency. However, MoE models are plagued by two fundamental challenges: 1) a load imbalance problem known as the``rich get richer" phenomenon, where a few experts are over-utilized, and 2) an expert homogeneity problem, where experts learn redundant representations, negating their purpose. Current solutions typically employ an auxiliary load-balancing loss that, while mitigating imbalance, often exacerbates homogeneity by enforcing uniform routing at the expense of specialization. To resolve this, we introduce the Eigen-Mixture-of-Experts (EMoE), a novel architecture that leverages a routing mechanism based on a learned orthonormal eigenbasis. EMoE projects input tokens onto this shared eigenbasis and routes them based on their alignment with the principal components of the feature space. This principled, geometric partitioning of data intrinsically promotes both balanced expert utilization and the development of diverse, specialized experts, all without the need for a conflicting auxiliary loss function. Our code is publicly available at https://github.com/Belis0811/EMoE.
MisSpans: Fine-Grained False Span Identification in Cross-Domain Fake News
Jan 08, 2026Abstract:Online misinformation is increasingly pervasive, yet most existing benchmarks and methods evaluate veracity at the level of whole claims or paragraphs using coarse binary labels, obscuring how true and false details often co-exist within single sentences. These simplifications also limit interpretability: global explanations cannot identify which specific segments are misleading or differentiate how a detail is false (e.g., distorted vs. fabricated). To address these gaps, we introduce MisSpans, the first multi-domain, human-annotated benchmark for span-level misinformation detection and analysis, consisting of paired real and fake news stories. MisSpans defines three complementary tasks: MisSpansIdentity for pinpointing false spans within sentences, MisSpansType for categorising false spans by misinformation type, and MisSpansExplanation for providing rationales grounded in identified spans. Together, these tasks enable fine-grained localisation, nuanced characterisation beyond true/false and actionable explanations. Expert annotators were guided by standardised guidelines and consistency checks, leading to high inter-annotator agreement. We evaluate 15 representative LLMs, including reasoning-enhanced and non-reasoning variants, under zero-shot and one-shot settings. Results reveal the challenging nature of fine-grained misinformation identification and analysis, and highlight the need for a deeper understanding of how performance may be influenced by multiple interacting factors, including model size and reasoning capabilities, along with domain-specific textual features. This project will be available at https://github.com/lzw108/MisSpans.
ERMoE: Eigen-Reparameterized Mixture-of-Experts for Stable Routing and Interpretable Specialization
Nov 14, 2025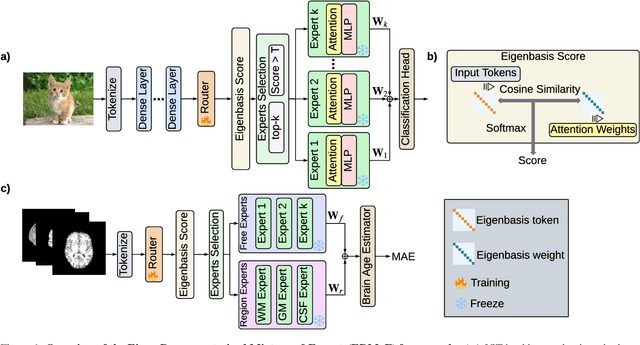



Abstract:Mixture-of-Experts (MoE) architectures expand model capacity by sparsely activating experts but face two core challenges: misalignment between router logits and each expert's internal structure leads to unstable routing and expert underutilization, and load imbalances create straggler bottlenecks. Standard solutions, such as auxiliary load-balancing losses, can reduce load disparities but often weaken expert specialization and hurt downstream performance. To address these issues, we propose ERMoE, a sparse MoE transformer that reparameterizes each expert in a learned orthonormal eigenbasis and replaces learned gating logits with an "Eigenbasis Score", defined as the cosine similarity between input features and an expert's basis. This content-aware routing ties token assignments directly to experts' representation spaces, stabilizing utilization and promoting interpretable specialization without sacrificing sparsity. Crucially, ERMoE removes the need for explicit balancing losses and avoids the interfering gradients they introduce. We show that ERMoE achieves state-of-the-art accuracy on ImageNet classification and cross-modal image-text retrieval benchmarks (e.g., COCO, Flickr30K), while naturally producing flatter expert load distributions. Moreover, a 3D MRI variant (ERMoE-ba) improves brain age prediction accuracy by more than 7\% and yields anatomically interpretable expert specializations. ERMoE thus introduces a new architectural principle for sparse expert models that directly addresses routing instabilities and enables improved performance with scalable, interpretable specialization.
ConspEmoLLM-v2: A robust and stable model to detect sentiment-transformed conspiracy theories
May 20, 2025Abstract:Despite the many benefits of large language models (LLMs), they can also cause harm, e.g., through automatic generation of misinformation, including conspiracy theories. Moreover, LLMs can also ''disguise'' conspiracy theories by altering characteristic textual features, e.g., by transforming their typically strong negative emotions into a more positive tone. Although several studies have proposed automated conspiracy theory detection methods, they are usually trained using human-authored text, whose features can vary from LLM-generated text. Furthermore, several conspiracy detection models, including the previously proposed ConspEmoLLM, rely heavily on the typical emotional features of human-authored conspiracy content. As such, intentionally disguised content may evade detection. To combat such issues, we firstly developed an augmented version of the ConDID conspiracy detection dataset, ConDID-v2, which supplements human-authored conspiracy tweets with versions rewritten by an LLM to reduce the negativity of their original sentiment. The quality of the rewritten tweets was verified by combining human and LLM-based assessment. We subsequently used ConDID-v2 to train ConspEmoLLM-v2, an enhanced version of ConspEmoLLM. Experimental results demonstrate that ConspEmoLLM-v2 retains or exceeds the performance of ConspEmoLLM on the original human-authored content in ConDID, and considerably outperforms both ConspEmoLLM and several other baselines when applied to sentiment-transformed tweets in ConDID-v2. The project will be available at https://github.com/lzw108/ConspEmoLLM.
Distributed Harmonization: Federated Clustered Batch Effect Adjustment and Generalization
May 23, 2024



Abstract:Independent and identically distributed (i.i.d.) data is essential to many data analysis and modeling techniques. In the medical domain, collecting data from multiple sites or institutions is a common strategy that guarantees sufficient clinical diversity, determined by the decentralized nature of medical data. However, data from various sites are easily biased by the local environment or facilities, thereby violating the i.i.d. rule. A common strategy is to harmonize the site bias while retaining important biological information. The ComBat is among the most popular harmonization approaches and has recently been extended to handle distributed sites. However, when faced with situations involving newly joined sites in training or evaluating data from unknown/unseen sites, ComBat lacks compatibility and requires retraining with data from all the sites. The retraining leads to significant computational and logistic overhead that is usually prohibitive. In this work, we develop a novel Cluster ComBat harmonization algorithm, which leverages cluster patterns of the data in different sites and greatly advances the usability of ComBat harmonization. We use extensive simulation and real medical imaging data from ADNI to demonstrate the superiority of the proposed approach.
Interpretable Spatio-Temporal Embedding for Brain Structural-Effective Network with Ordinary Differential Equation
May 21, 2024



Abstract:The MRI-derived brain network serves as a pivotal instrument in elucidating both the structural and functional aspects of the brain, encompassing the ramifications of diseases and developmental processes. However, prevailing methodologies, often focusing on synchronous BOLD signals from functional MRI (fMRI), may not capture directional influences among brain regions and rarely tackle temporal functional dynamics. In this study, we first construct the brain-effective network via the dynamic causal model. Subsequently, we introduce an interpretable graph learning framework termed Spatio-Temporal Embedding ODE (STE-ODE). This framework incorporates specifically designed directed node embedding layers, aiming at capturing the dynamic interplay between structural and effective networks via an ordinary differential equation (ODE) model, which characterizes spatial-temporal brain dynamics. Our framework is validated on several clinical phenotype prediction tasks using two independent publicly available datasets (HCP and OASIS). The experimental results clearly demonstrate the advantages of our model compared to several state-of-the-art methods.
ConspEmoLLM: Conspiracy Theory Detection Using an Emotion-Based Large Language Model
Mar 11, 2024



Abstract:The internet has brought both benefits and harms to society. A prime example of the latter is misinformation, including conspiracy theories, which flood the web. Recent advances in natural language processing, particularly the emergence of large language models (LLMs), have improved the prospects of accurate misinformation detection. However, most LLM-based approaches to conspiracy theory detection focus only on binary classification and fail to account for the important relationship between misinformation and affective features (i.e., sentiment and emotions). Driven by a comprehensive analysis of conspiracy text that reveals its distinctive affective features, we propose ConspEmoLLM, the first open-source LLM that integrates affective information and is able to perform diverse tasks relating to conspiracy theories. These tasks include not only conspiracy theory detection, but also classification of theory type and detection of related discussion (e.g., opinions towards theories). ConspEmoLLM is fine-tuned based on an emotion-oriented LLM using our novel ConDID dataset, which includes five tasks to support LLM instruction tuning and evaluation. We demonstrate that when applied to these tasks, ConspEmoLLM largely outperforms several open-source general domain LLMs and ChatGPT, as well as an LLM that has been fine-tuned using ConDID, but which does not use affective features. This project will be released on https://github.com/lzw108/ConspEmoLLM/.
Emotion Detection for Misinformation: A Review
Nov 01, 2023



Abstract:With the advent of social media, an increasing number of netizens are sharing and reading posts and news online. However, the huge volumes of misinformation (e.g., fake news and rumors) that flood the internet can adversely affect people's lives, and have resulted in the emergence of rumor and fake news detection as a hot research topic. The emotions and sentiments of netizens, as expressed in social media posts and news, constitute important factors that can help to distinguish fake news from genuine news and to understand the spread of rumors. This article comprehensively reviews emotion-based methods for misinformation detection. We begin by explaining the strong links between emotions and misinformation. We subsequently provide a detailed analysis of a range of misinformation detection methods that employ a variety of emotion, sentiment and stance-based features, and describe their strengths and weaknesses. Finally, we discuss a number of ongoing challenges in emotion-based misinformation detection based on large language models and suggest future research directions, including data collection (multi-platform, multilingual), annotation, benchmark, multimodality, and interpretability.
Towards Sparsified Federated Neuroimaging Models via Weight Pruning
Aug 24, 2022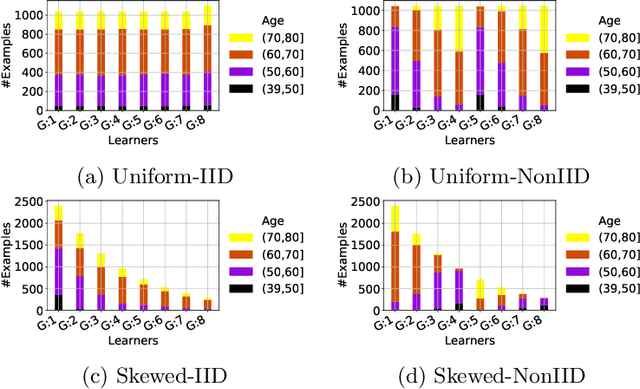
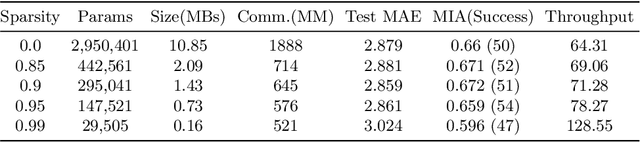
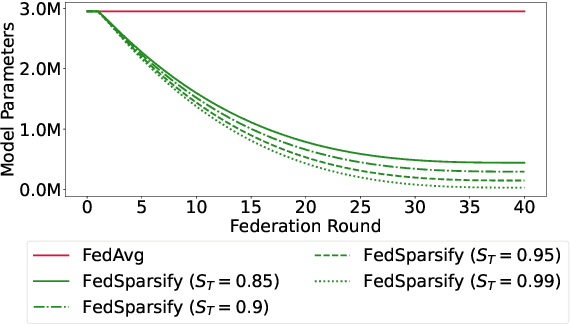
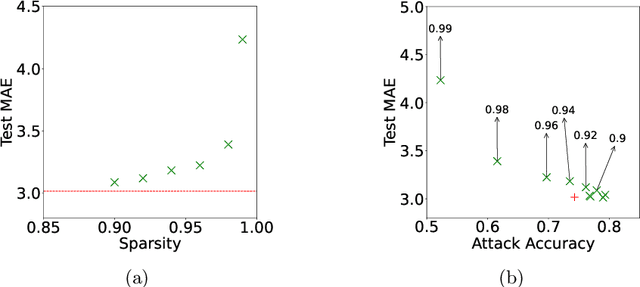
Abstract:Federated training of large deep neural networks can often be restrictive due to the increasing costs of communicating the updates with increasing model sizes. Various model pruning techniques have been designed in centralized settings to reduce inference times. Combining centralized pruning techniques with federated training seems intuitive for reducing communication costs -- by pruning the model parameters right before the communication step. Moreover, such a progressive model pruning approach during training can also reduce training times/costs. To this end, we propose FedSparsify, which performs model pruning during federated training. In our experiments in centralized and federated settings on the brain age prediction task (estimating a person's age from their brain MRI), we demonstrate that models can be pruned up to 95% sparsity without affecting performance even in challenging federated learning environments with highly heterogeneous data distributions. One surprising benefit of model pruning is improved model privacy. We demonstrate that models with high sparsity are less susceptible to membership inference attacks, a type of privacy attack.
Functional2Structural: Cross-Modality Brain Networks Representation Learning
May 06, 2022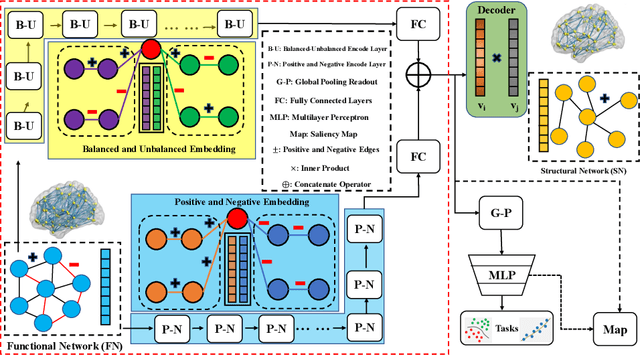
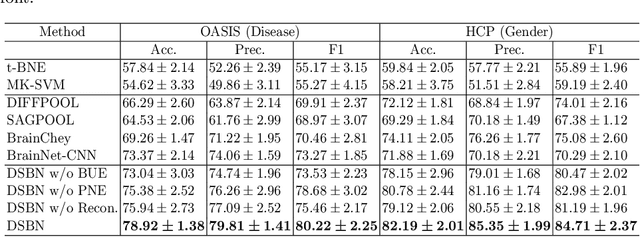
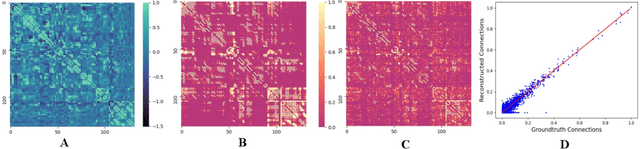
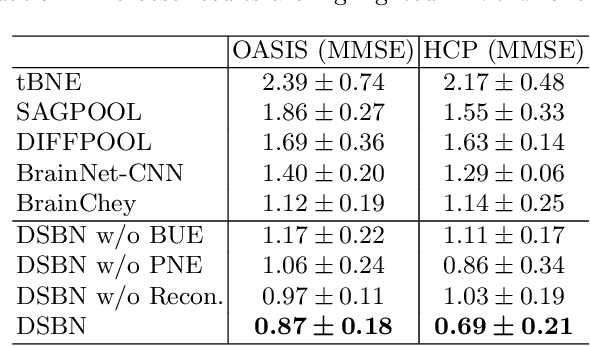
Abstract:MRI-based modeling of brain networks has been widely used to understand functional and structural interactions and connections among brain regions, and factors that affect them, such as brain development and disease. Graph mining on brain networks may facilitate the discovery of novel biomarkers for clinical phenotypes and neurodegenerative diseases. Since brain networks derived from functional and structural MRI describe the brain topology from different perspectives, exploring a representation that combines these cross-modality brain networks is non-trivial. Most current studies aim to extract a fused representation of the two types of brain network by projecting the structural network to the functional counterpart. Since the functional network is dynamic and the structural network is static, mapping a static object to a dynamic object is suboptimal. However, mapping in the opposite direction is not feasible due to the non-negativity requirement of current graph learning techniques. Here, we propose a novel graph learning framework, known as Deep Signed Brain Networks (DSBN), with a signed graph encoder that, from an opposite perspective, learns the cross-modality representations by projecting the functional network to the structural counterpart. We validate our framework on clinical phenotype and neurodegenerative disease prediction tasks using two independent, publicly available datasets (HCP and OASIS). The experimental results clearly demonstrate the advantages of our model compared to several state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge